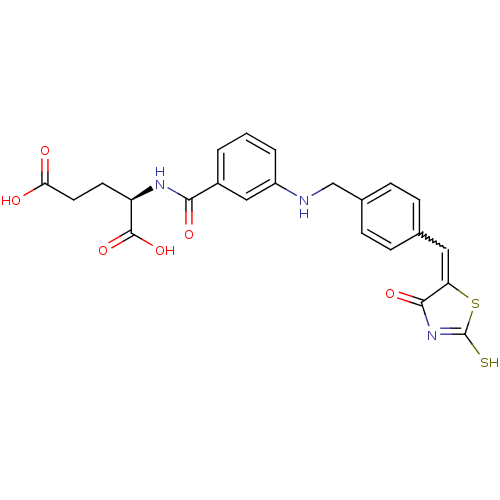
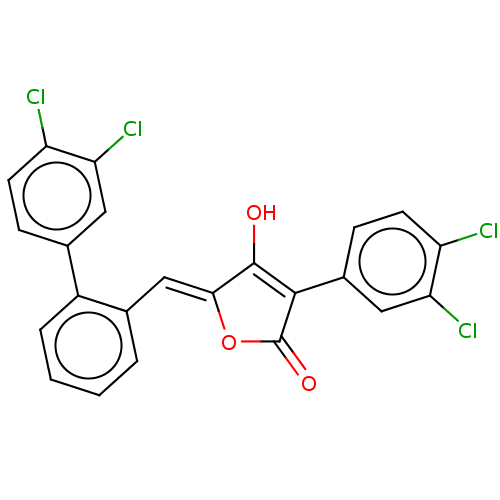
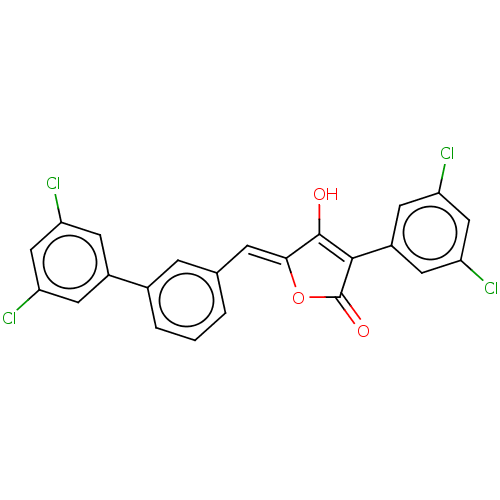
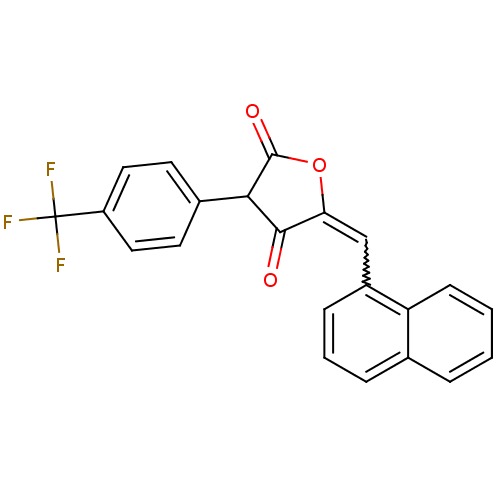
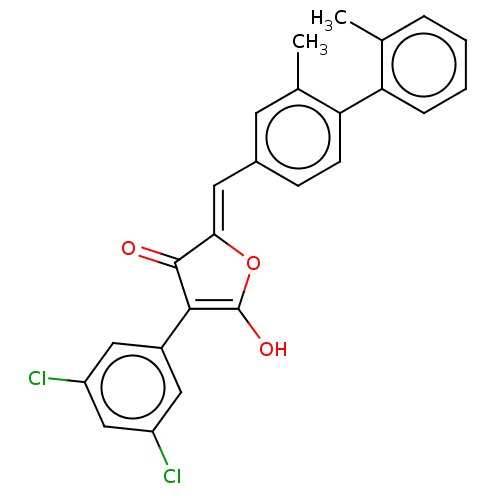
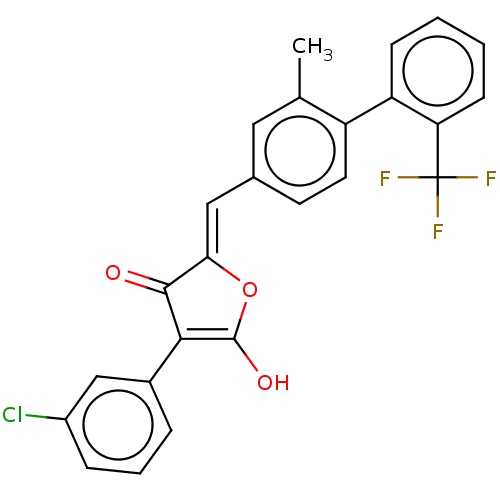
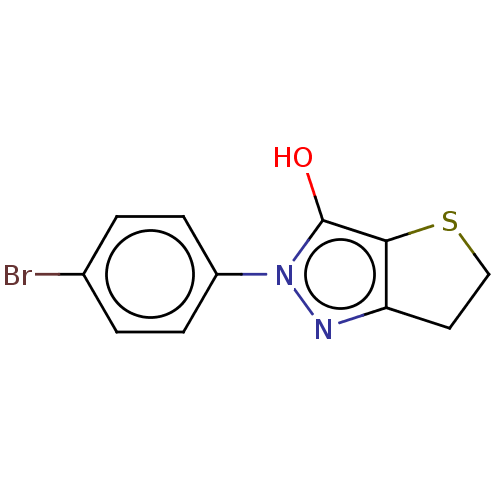
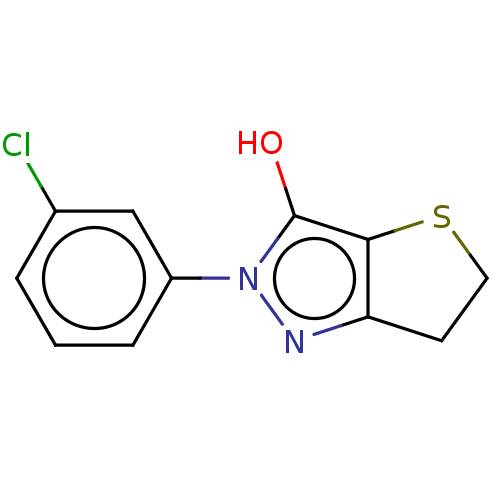
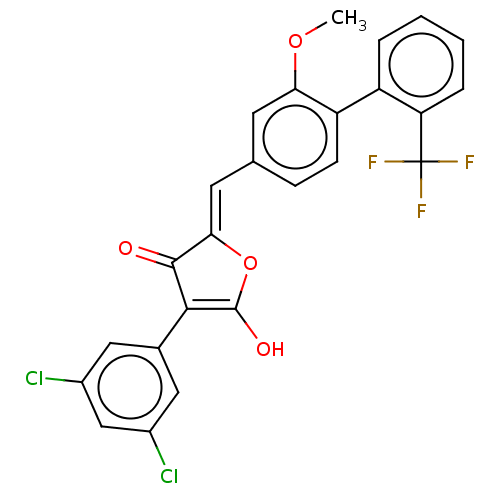
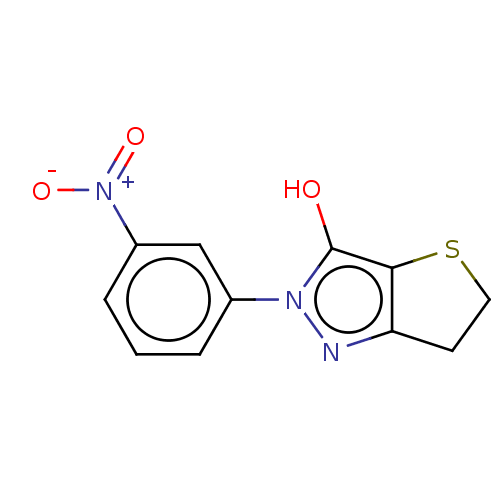
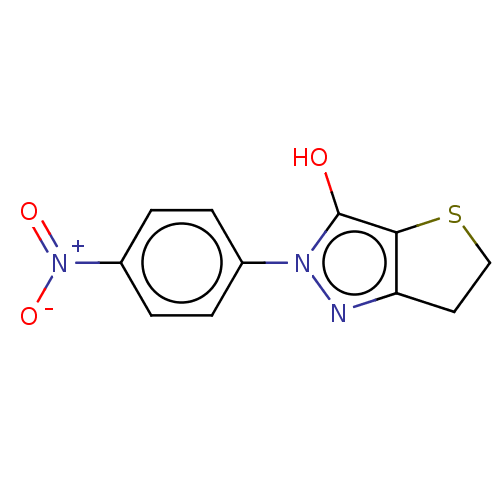
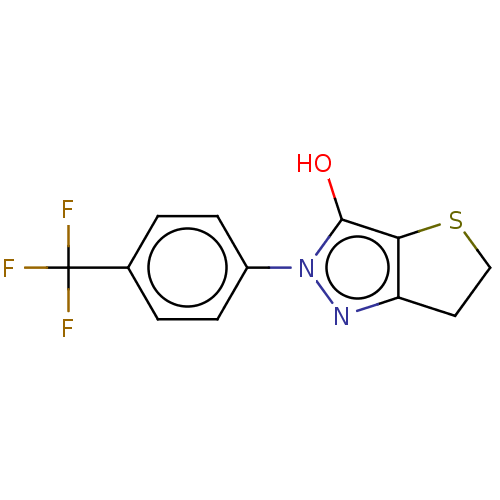
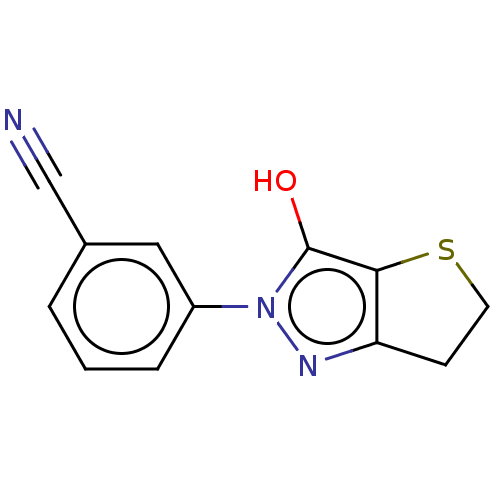
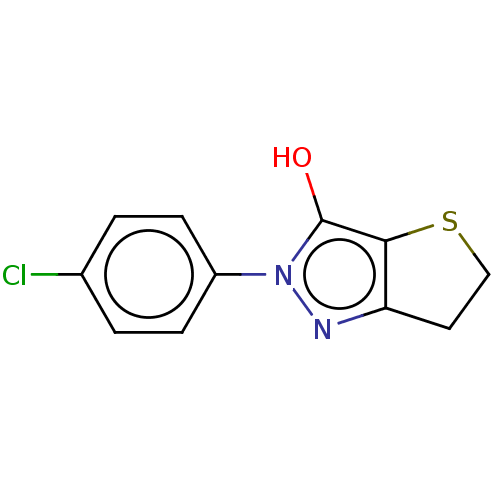
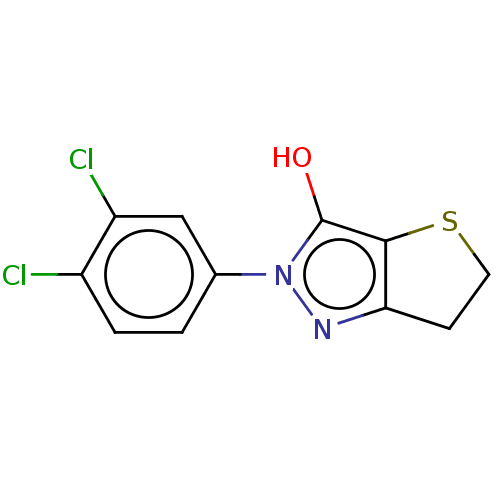
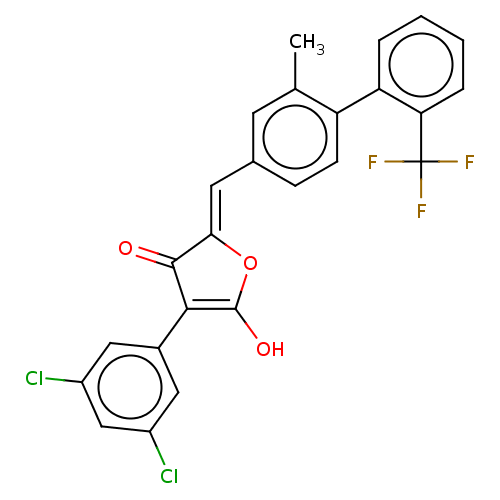
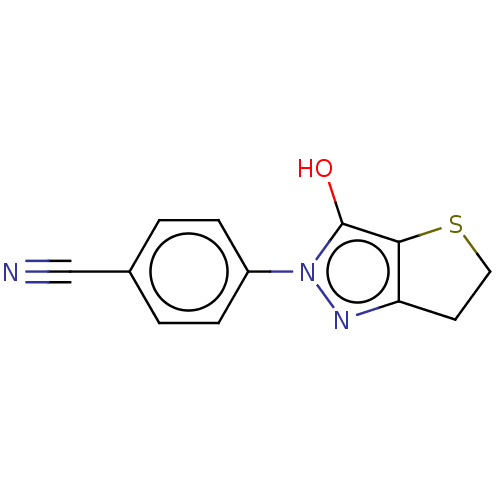
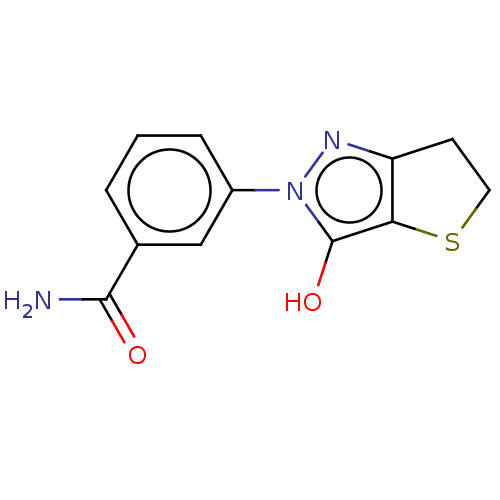
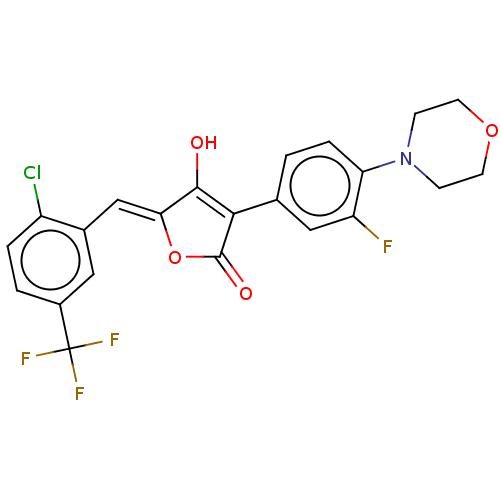
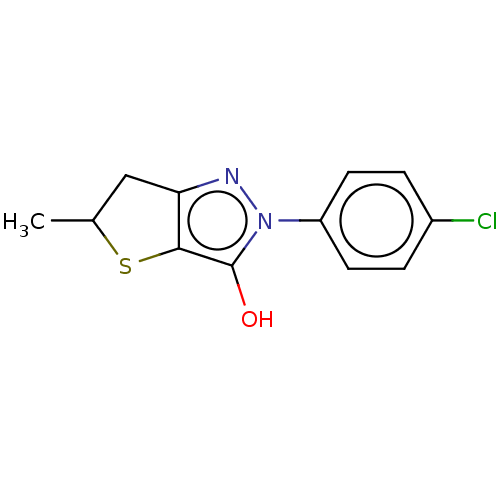
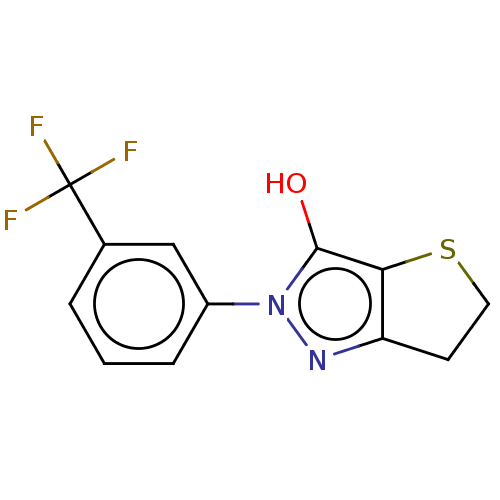
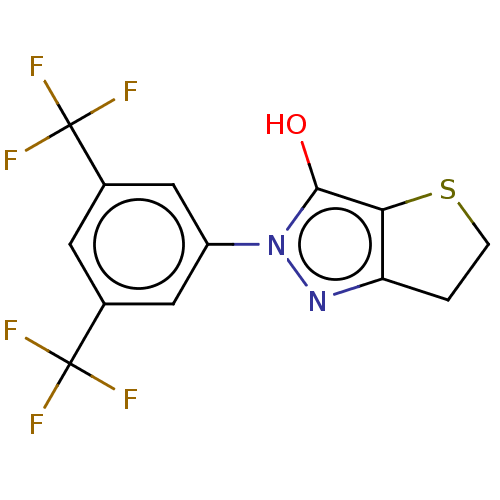
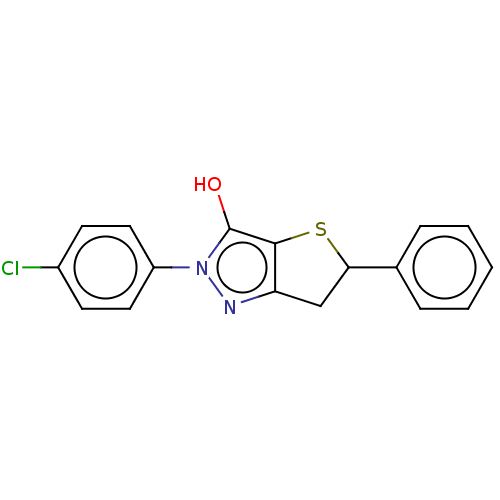
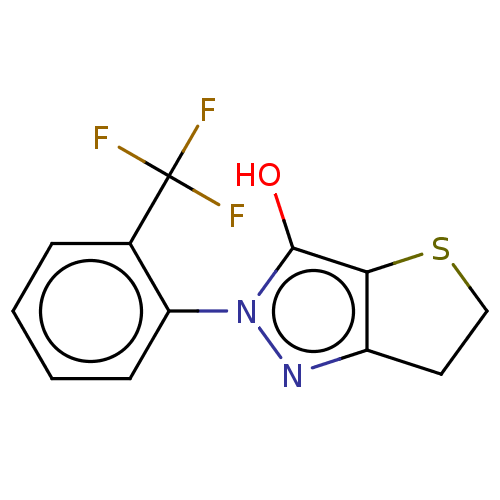
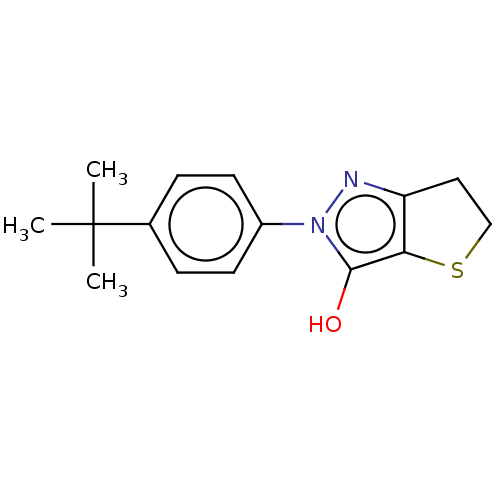
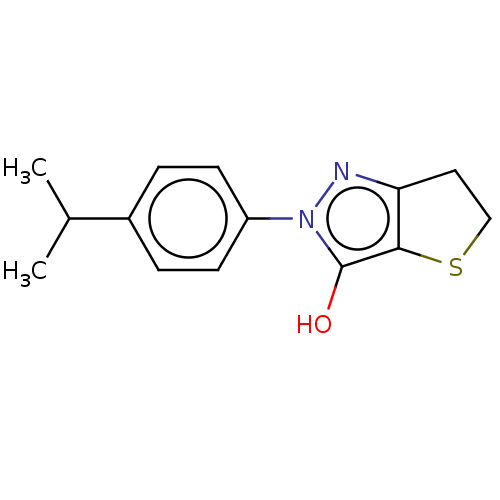
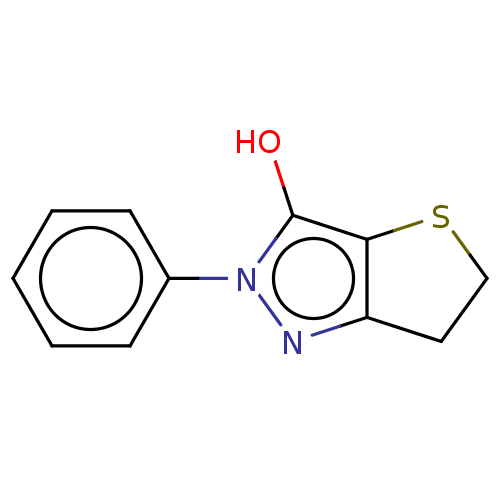
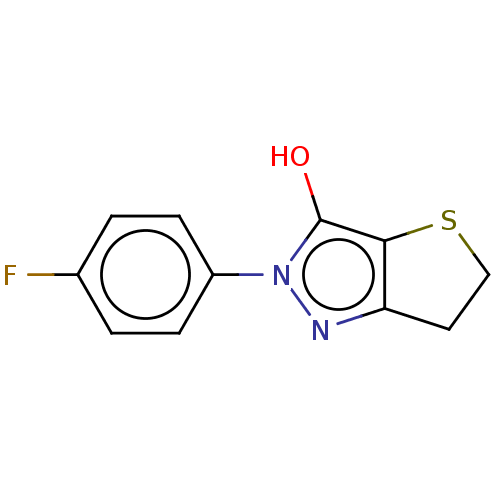
Affinity DataIC50: 1.04E+4nMAssay Description:Inhibitory activity against MurD in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+4nMAssay Description:Inhibitory activity against MurD in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 2.05E+4nMAssay Description:Inhibitory activity against MurD in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 2.62E+4nMAssay Description:Inhibitory activity against MurD in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 3.02E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 3.28E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 3.35E+4nMAssay Description:Inhibitory activity against MurD in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 3.45E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.19E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.52E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.74E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.87E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.88E+4nMAssay Description:Inhibitory activity against MurD in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 4.93E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.97E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 5.32E+4nMAssay Description:Inhibitory activity against MurD in Staphylococcus aureusMore data for this Ligand-Target Pair
Affinity DataIC50: 6.37E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 6.63E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 7.05E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 7.60E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 8.73E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.11E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.60E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.62E+4nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+5nMAssay Description:In vitro inhibitory activity against Staphylococcus aureus UDP-N-acetylmuramoyl-L-alanyl-D-glutamate synthetaseMore data for this Ligand-Target Pair