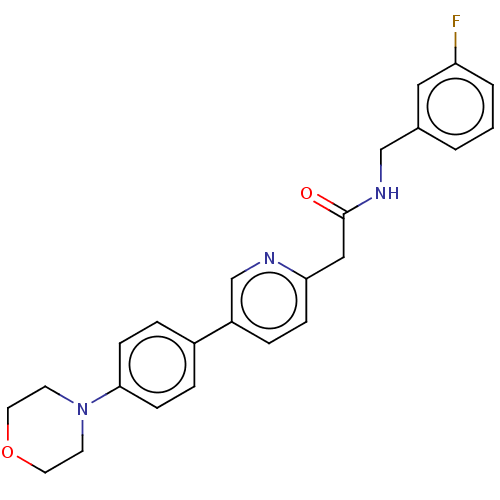
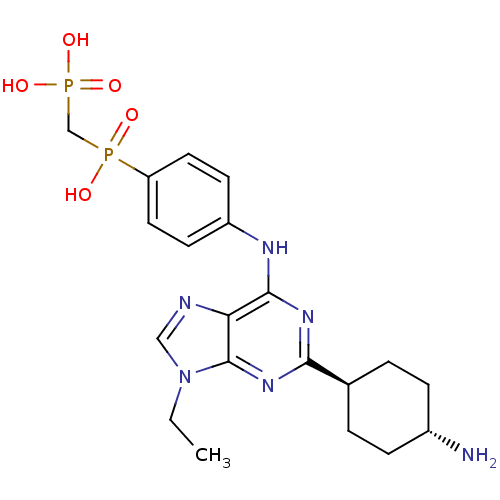
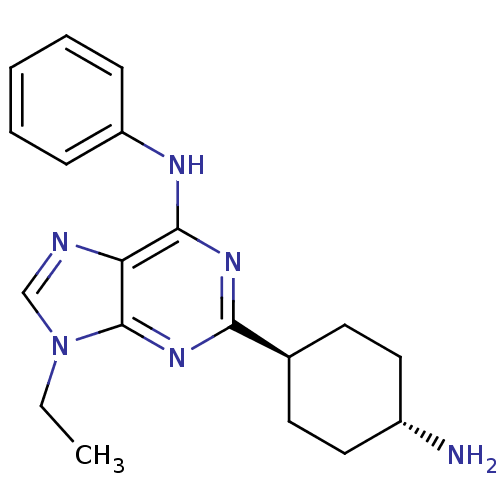
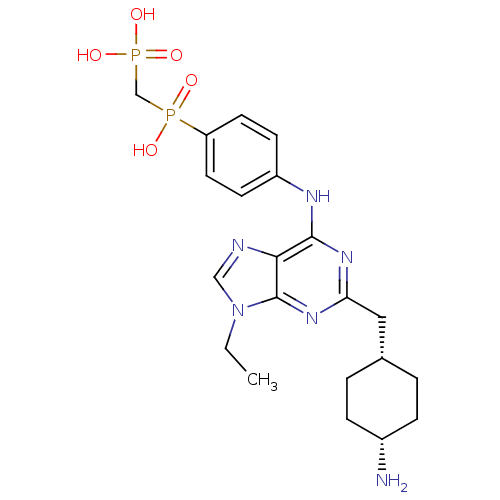
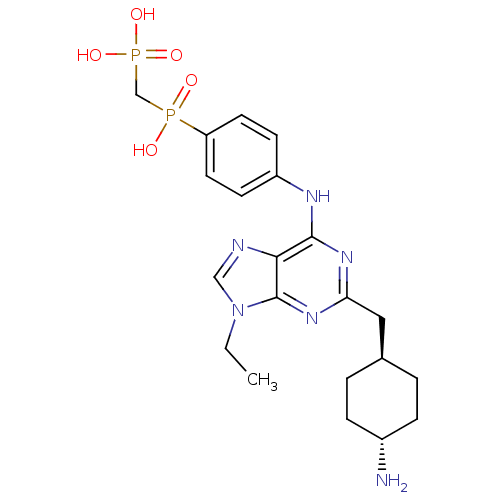
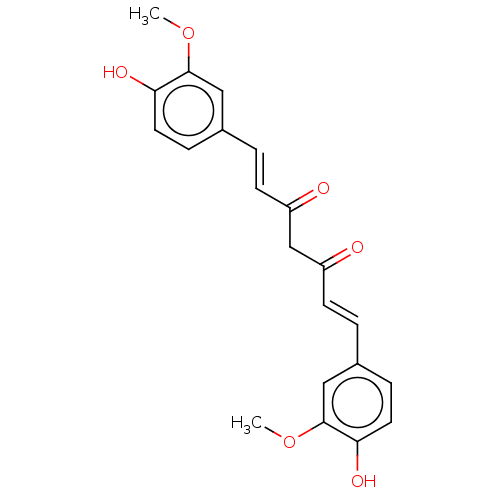
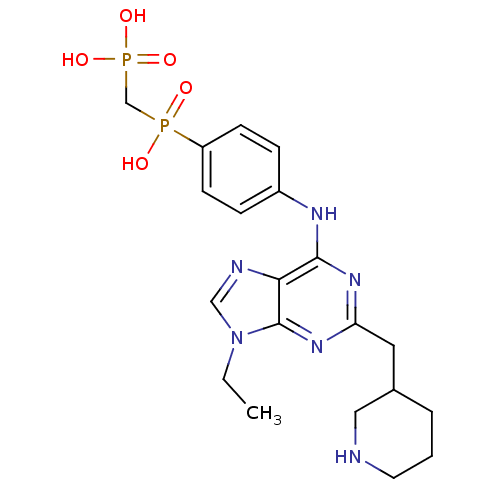
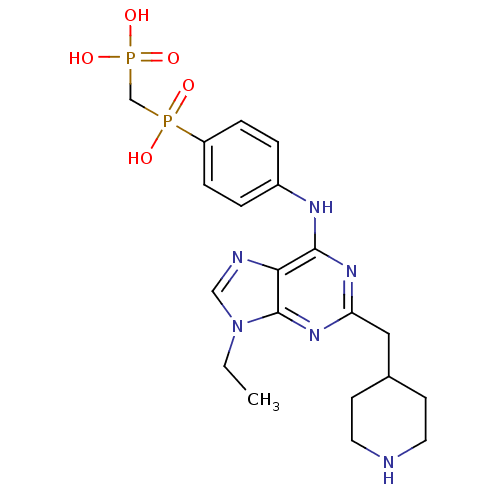
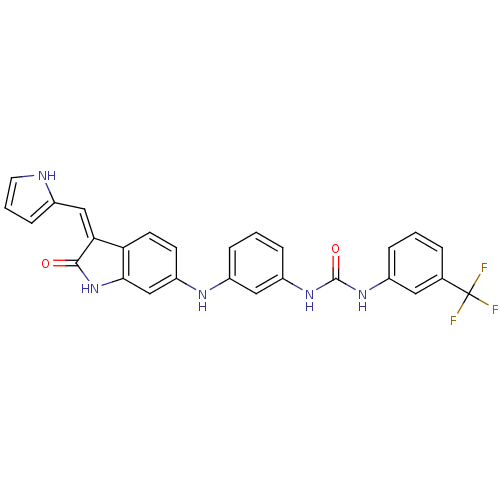
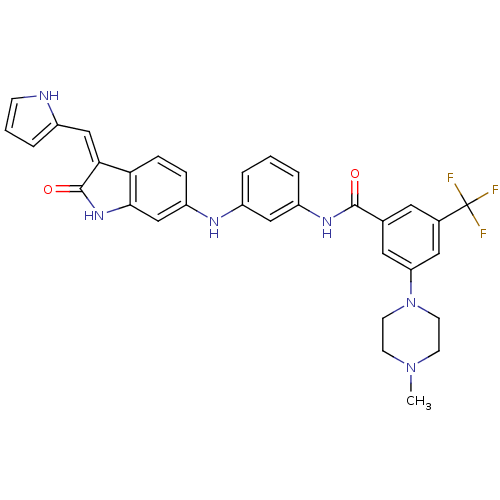
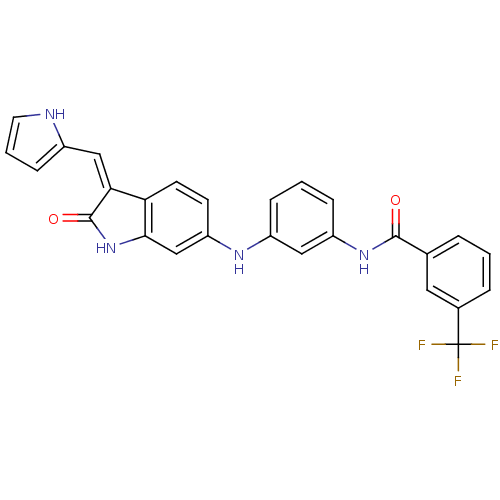
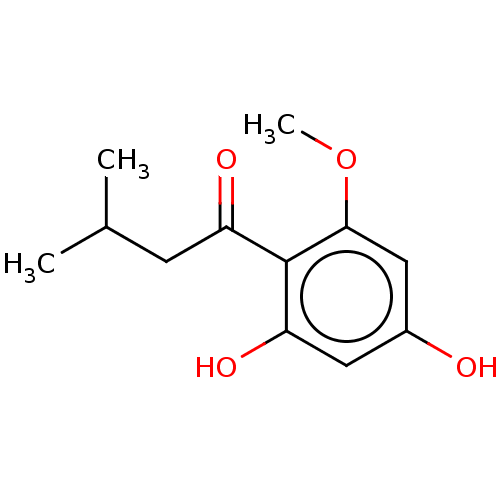
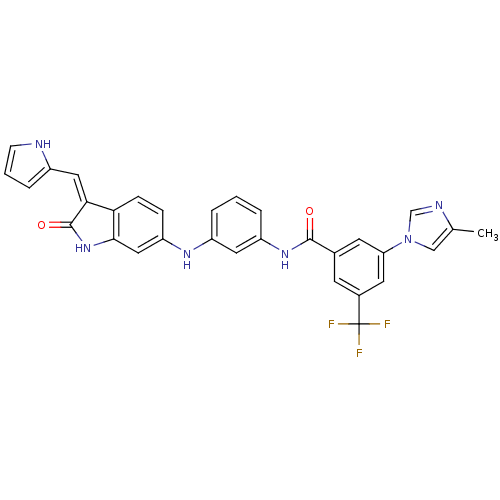
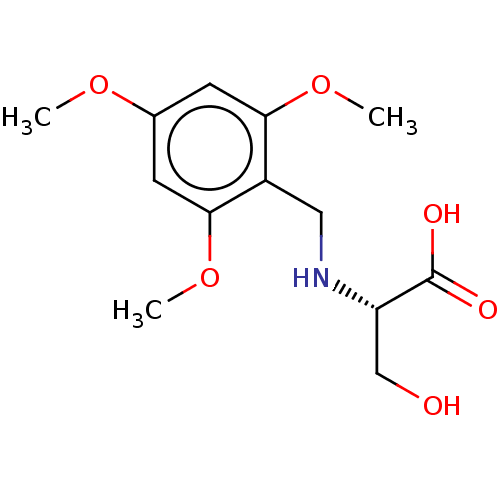
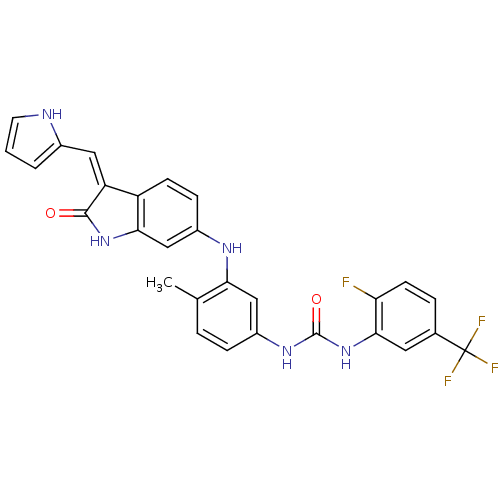
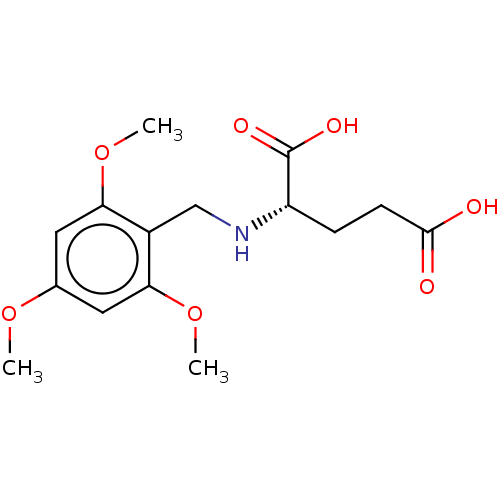
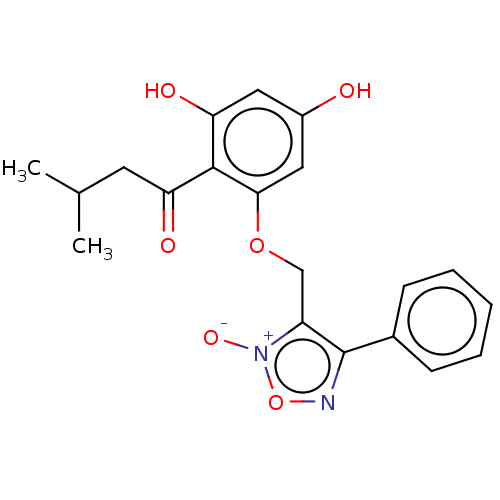
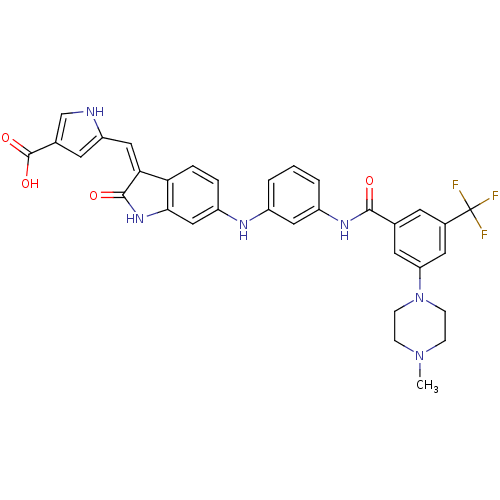
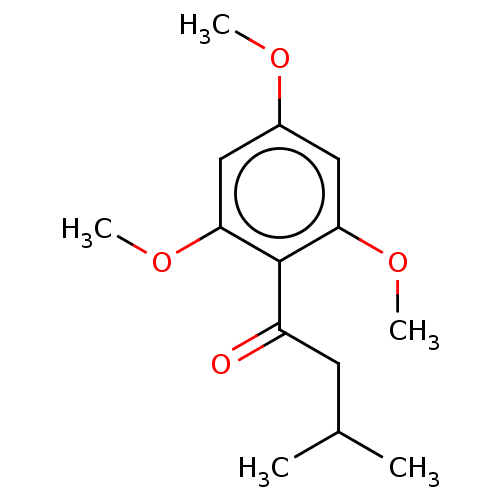
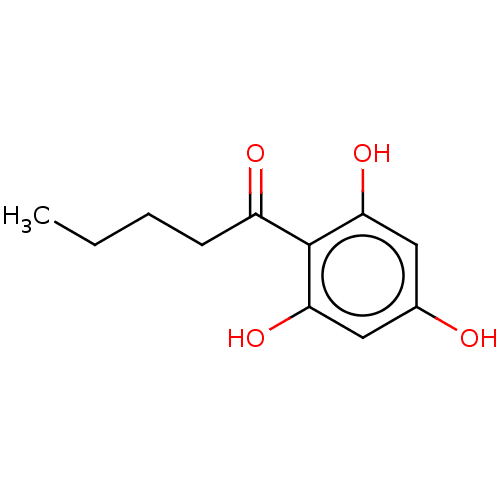
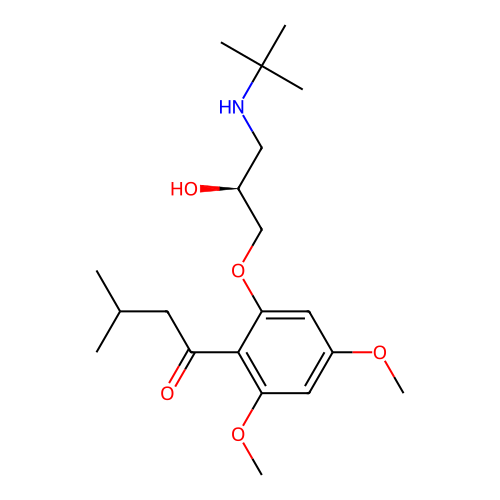
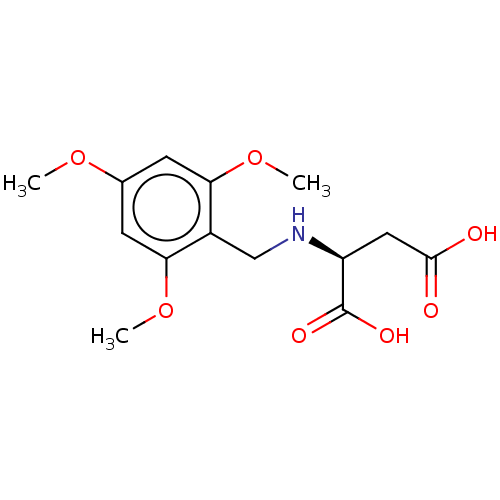
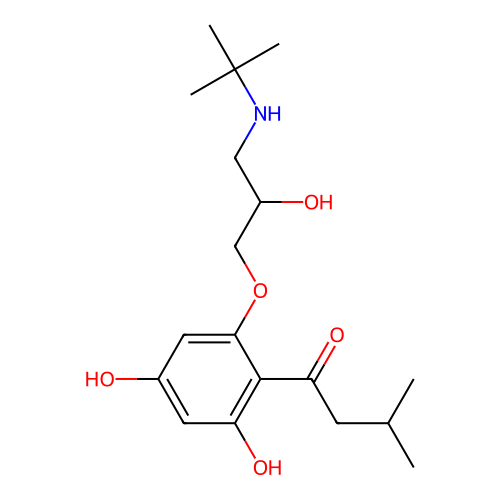
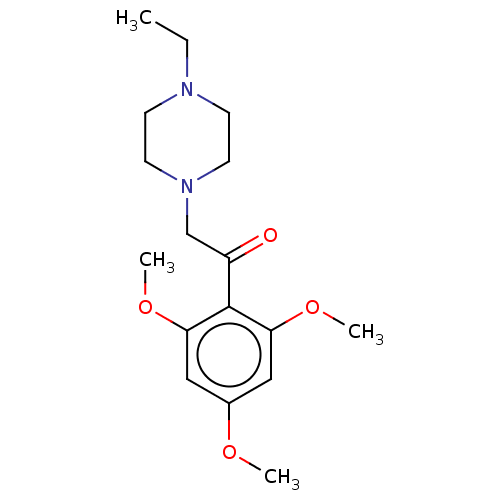
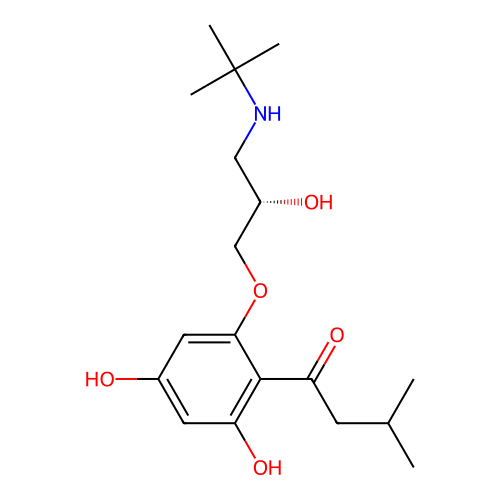
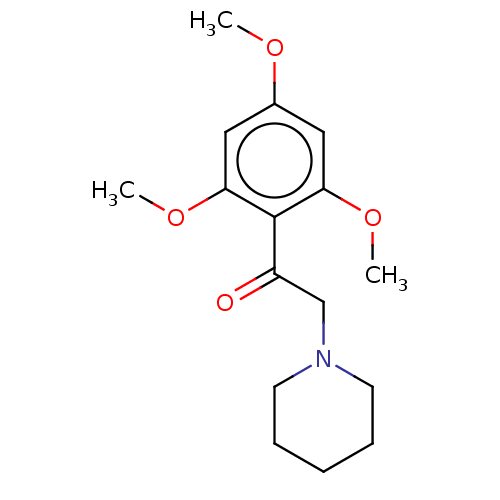
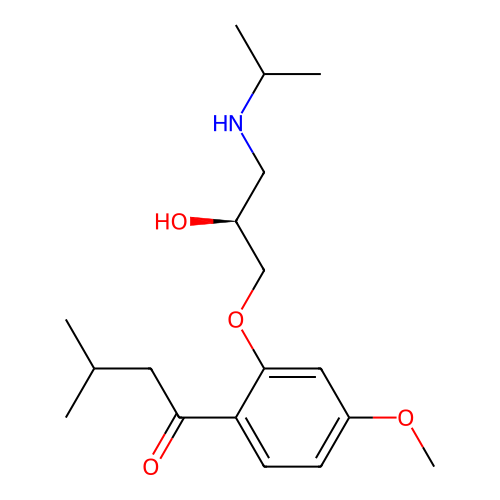
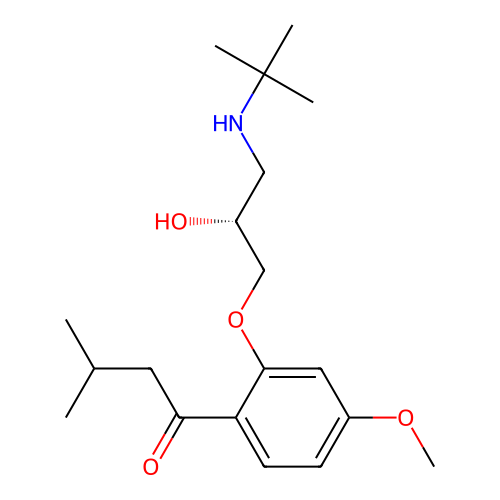
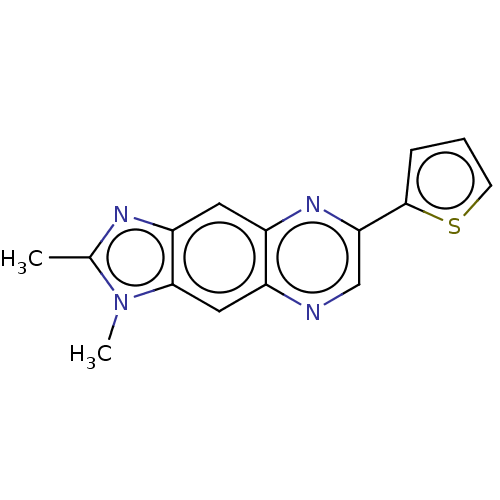
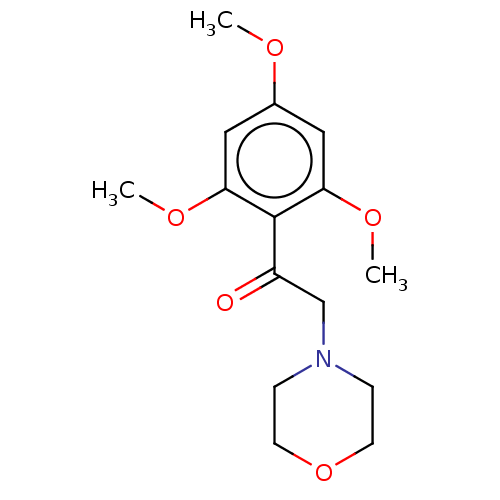
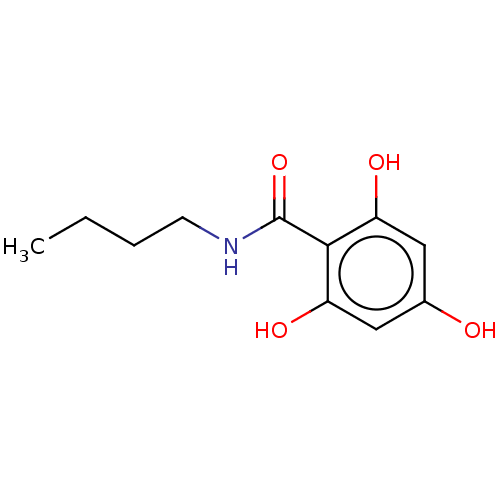
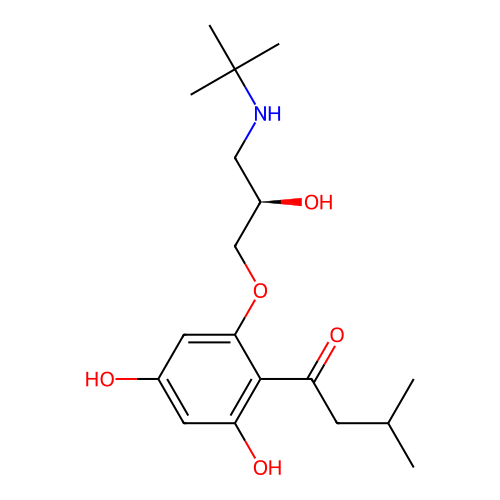
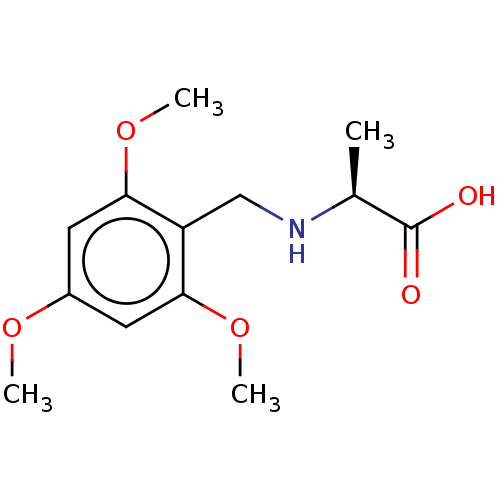
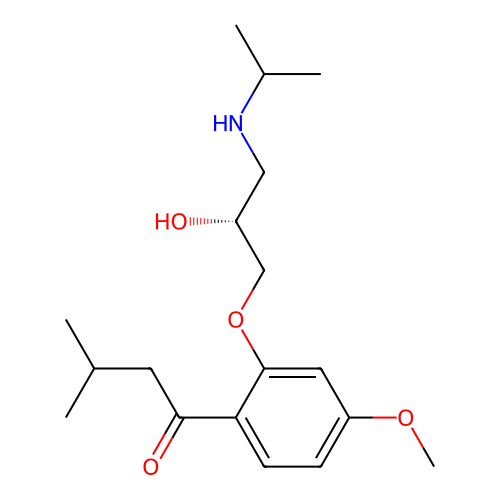
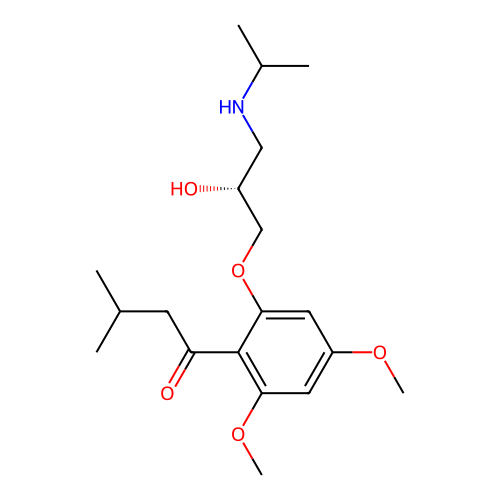
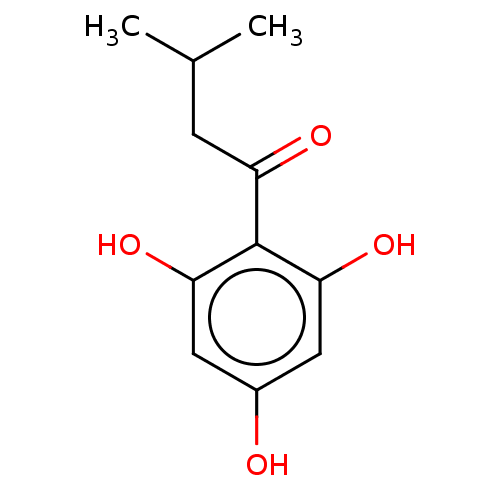
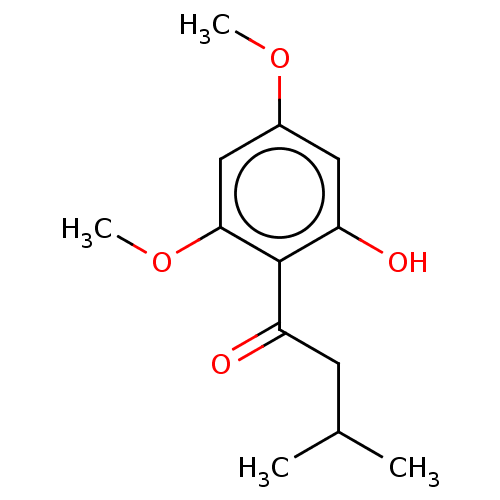
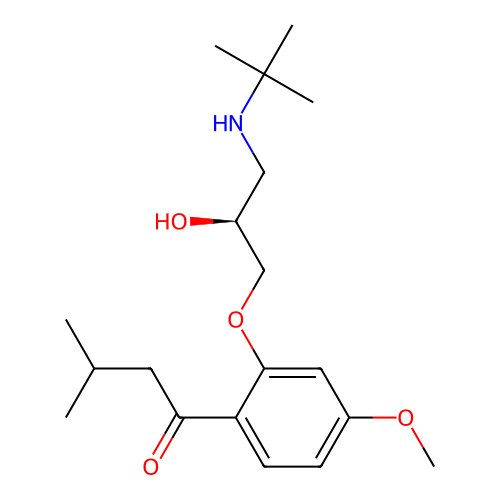
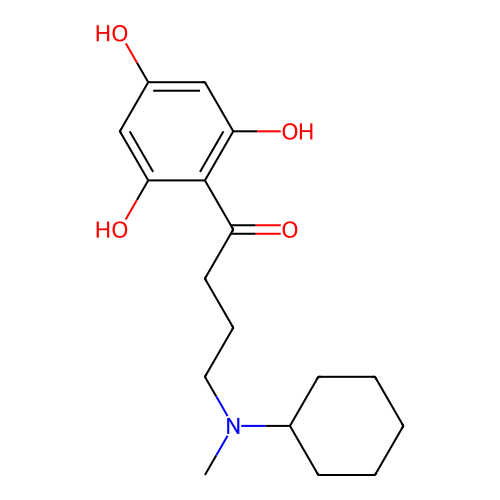
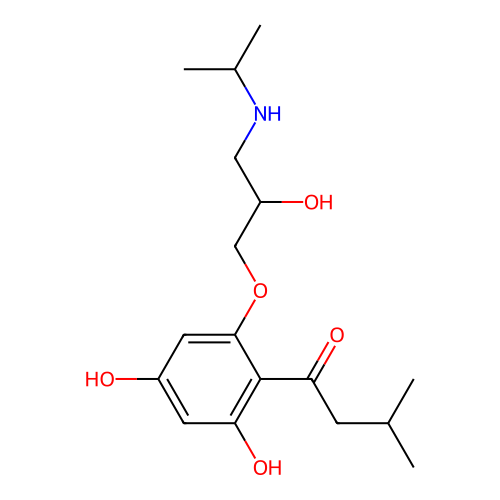
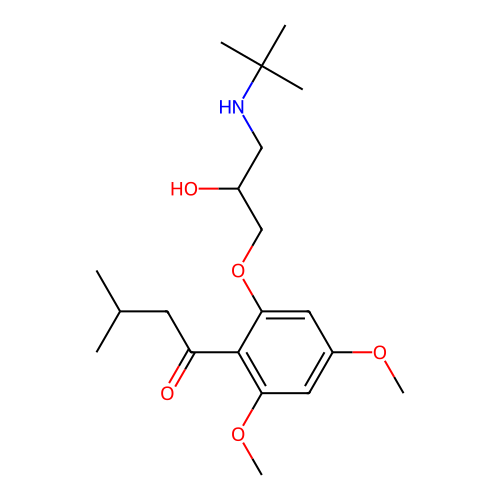
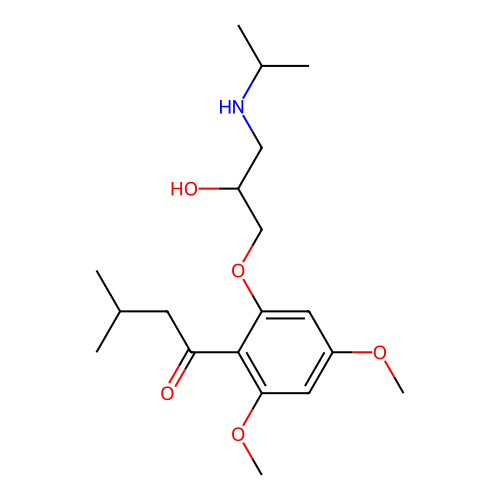
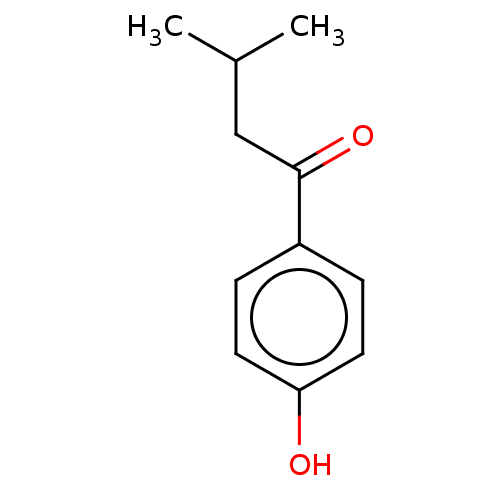
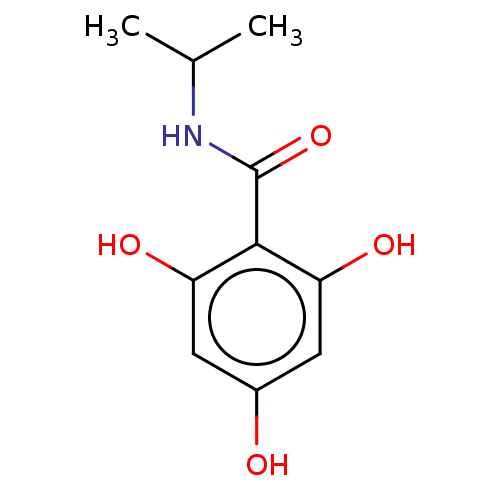
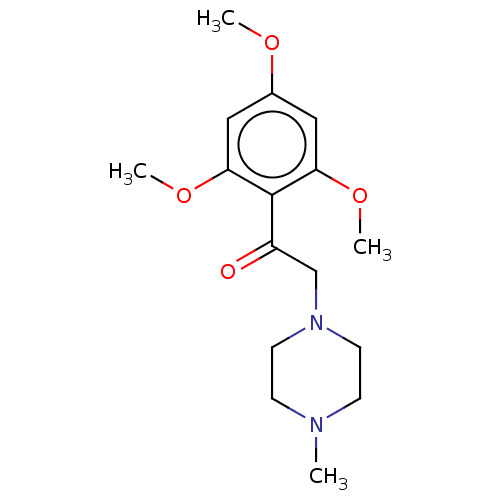
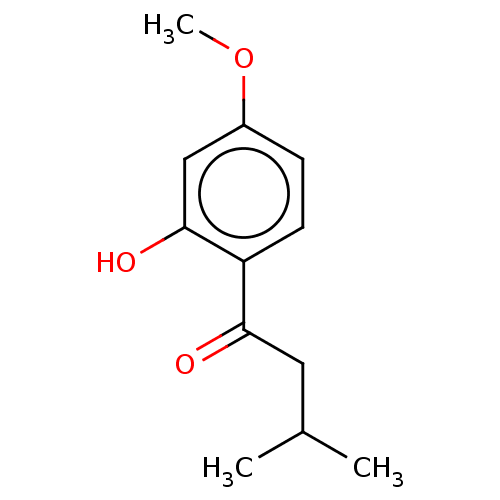
Affinity DataIC50: 60nMAssay Description:Inhibition of Src autophosphorylation in mouse GL261 cellsMore data for this Ligand-Target Pair
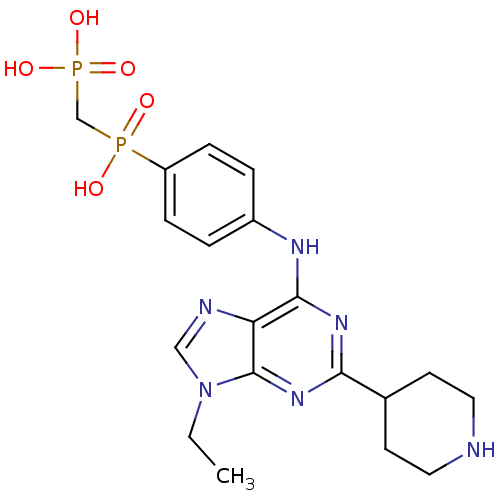
Affinity DataIC50: 68nMAssay Description:2,6,9-Trisubstituted purines were evaluated for their src kinase inhibitory activities using an ELISA assay.More data for this Ligand-Target Pair
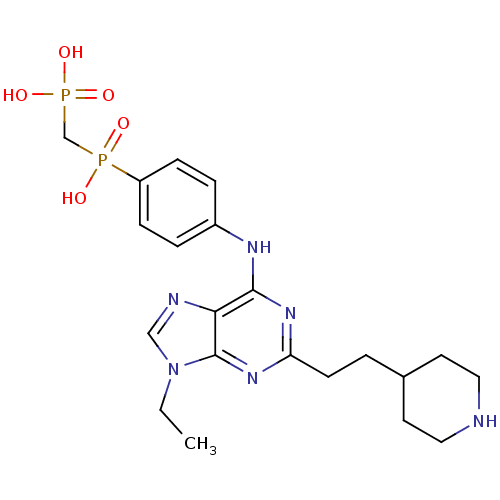
Affinity DataIC50: 73nMAssay Description:2,6,9-Trisubstituted purines were evaluated for their src kinase inhibitory activities using an ELISA assay.More data for this Ligand-Target Pair
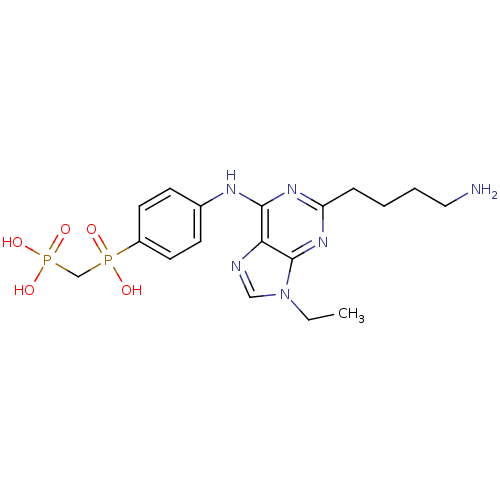
Affinity DataIC50: 101nMAssay Description:2,6,9-Trisubstituted purines were evaluated for their src kinase inhibitory activities using an ELISA assay.More data for this Ligand-Target Pair
Affinity DataIC50: 205nMAssay Description:2,6,9-Trisubstituted purines were evaluated for their src kinase inhibitory activities using an ELISA assay.More data for this Ligand-Target Pair
Affinity DataIC50: 273nMAssay Description:2,6,9-Trisubstituted purines were evaluated for their src kinase inhibitory activities using an ELISA assay.More data for this Ligand-Target Pair
Affinity DataIC50: 665nMAssay Description:2,6,9-Trisubstituted purines were evaluated for their src kinase inhibitory activities using an ELISA assay.More data for this Ligand-Target Pair
Affinity DataIC50: 1.01E+3nMAssay Description:2,6,9-Trisubstituted purines were evaluated for their src kinase inhibitory activities using an ELISA assay.More data for this Ligand-Target Pair
Affinity DataIC50: 2.11E+3nMAssay Description:2,6,9-Trisubstituted purines were evaluated for their src kinase inhibitory activities using an ELISA assay.More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:2,6,9-Trisubstituted purines were evaluated for their src kinase inhibitory activities using an ELISA assay.More data for this Ligand-Target Pair
Affinity DataIC50: 2.31E+3nMAssay Description:2,6,9-Trisubstituted purines were evaluated for their src kinase inhibitory activities using an ELISA assay.More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of Tel-fused SRC in mouse BA/F3 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+3nMAssay Description:Inhibition of Tel-fused SRC in mouse BA/F3 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+3nMAssay Description:Inhibition of Tel-fused SRC in mouse BA/F3 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+3nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+3nMAssay Description:Inhibition of Tel-fused SRC in mouse BA/F3 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 6.60E+3nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.30E+3nMAssay Description:Inhibition of Tel-fused SRC in mouse BA/F3 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 8.40E+3nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.50E+3nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Tel-fused SRC in mouse BA/F3 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.11E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.12E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.12E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.44E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.17E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.27E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.41E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.49E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.56E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.82E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of SRC in transformed mouse NIH3T3 cells by SDS-PAGE based immunoblot assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.24E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.71E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.73E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.91E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.09E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.13E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.06E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.15E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.38E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.48E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.40E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.50E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.48E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.07E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.26E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.53E+4nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.13E+5nMAssay Description:Inhibition of Src in mouse BV2 cells assessed as suppression of LPS-induced NO release preincubated for 1 hr followed by LPS addition measured after ...More data for this Ligand-Target Pair